-
use OpenFOAM in docker
OS: windows 10
Install docker for windows
site: https://www.docker.com/
download: Docker for Windows Installer.exe
edition: most recent stable, for me, it is Docker version 17.09.0, community edition
installation instructions: https://docs.docker.com/docker-for-windows/
test
after completion of the installation, run
Docker for Windows. It will cost some time to start thedocker engine.right click "Start" button, choose "Windows Powershell"
docker --version ## Docker version 17.09.0-ce, build afdb6d4 docker-compose --version ## docker-compose version 1.16.1, build 6d1ac219 docker-machine --version ## docker-machine.exe version 0.12.2, build 9371605 docker run hello-world ## ## Hello from Docker! ## This message shows that your installation appears to be working correctly. ## ## To generate this message, Docker took the following steps: ## 1. The Docker client contacted the Docker daemon. ## 2. The Docker daemon pulled the "hello-world" image from the Docker Hub. ## (amd64) ## 3. The Docker daemon created a new container from that image which runs the ## executable that produces the output you are currently reading. ## 4. The Docker daemon streamed that output to the Docker client, which sent it ## to your terminal. ## ## To try something more ambitious, you can run an Ubuntu container with: ## $ docker run -it ubuntu bash ## ## Share images, automate workflows, and more with a free Docker ID: ## https://cloud.docker.com/ ## ## For more examples and ideas, visit: ## https://docs.docker.com/engine/userguide/ docker ps -a ## CONTAINER ID IMAGE COMMAND CREATED STATUS PORTS NAMES ## 32a621d46d34 hello-world "/hello" 24 seconds ago Exited (0) 23 seconds ago clever_agnesi docker rm 32a621d46d34 #change it to your container ID ## 32a621d46d34 docker ps -a ## CONTAINER ID IMAGE COMMAND CREATED STATUS PORTS NAMESdownload OpenFOAM image
right click "Start" button, choose "Windows Powershell"
docker pull openfoamplus/of_v1706_centos73 # OF+ # or docker pull openfoam/openfoam5-paraview54 # OF5 # list images docker image lsrun OpenFOAM
Right click on the "Docker" icon in the
system tray, choose "settings"Click "Shared drives"
choose any drive you want to share, in my case, I choose C, then click "apply"
you may need to input credentials
use
openfoam+1706as example.press "WIN+R", input
cmdand enterrem test docker run --rm -v c:/Users:/data alpine ls /data rem ONLY `C:/Users` and its subfolders can be mounted docker run ^ -i -t ^ --name myOFplus_1706 ^ -v c:/Users/dic17007/OpenFOAM:/OF ^ openfoamplus/of_v1706_centos73 ^ bash rem press `Ctrl+p`, `Ctrl+q` to return without stop the container docker attach myOFplus_1706 rem press `Ctrl+c` or input `exit` to return and stop the containerNote: you are
rootuser in docker and the password isofuser2017. Referencesetup the environment:
alias of1706="HOME=/OF source $DOCKER_OPENFOAM_PATH" # add to `~/.bashrc` # echo 'alias of1706="HOME=/OF source $DOCKER_OPENFOAM_PATH"'>>~/.bashrcrun testcase
of1706 #activate openfoam mkdir -p $FOAM_RUN run pwd #/OF/OpenFOAM/-v1706/run cp $FOAM_TUTORIALS/incompressible/icoFoam/cavity/cavity -r . cd cavity foamJob -screen blockMesh foamJob -screen icoFoam touch a.foam ## use Ctrl+P, Ctrl+Q to return to `cmd` ## use `docker attach myOFplus_1706`return to windows, use paraview to do post-process.
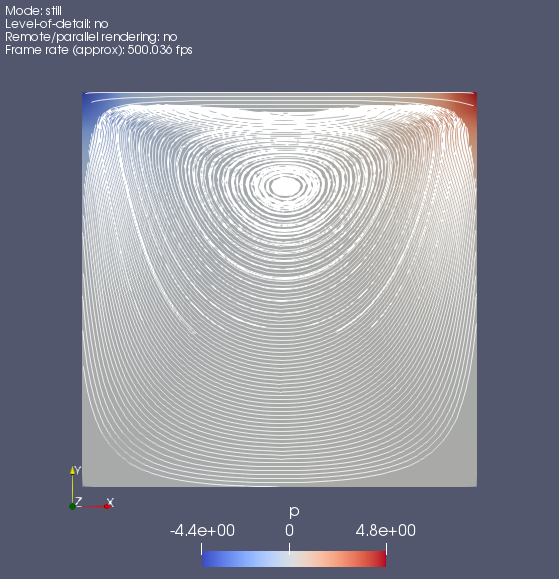
modify OpenFOAM solver
replication
run cd .. mkdir -p applications/solvers cd applications/solvers # I put my solver here. pwd # /OF/OpenFOAM/-v1706/applications/solvers cp $FOAM_SOLVERS/incompressible/icoFoam -r . mv icoFoam myIcoFoam cd myIcoFoam mv icoFoam.C myIcoFoam.C sed -i s/icoFoam/myIcoFoam/g myIcoFoam.C sed -i s/icoFoam/myIcoFoam/g Make/files sed -i s/FOAM_APPBIN/FOAM_USER_APPBIN Make/filesmake
wmaketest
run cd cavity which myIcoFoam foamJob -screen myIcoFoammodification
I am trying to output the matrix in COO format ( Coordinate Format). It will be consists of three parts:
- AA: non-zero values
- JR: row index
- JC: column index
According to the definition of
Foam::lduMatrix::Amul()function, there are 4 part of scalar matrix:-
diagonal terms: JC, JR=1 ... nCells, AA =
matrix.diag(); -
upper terms: JC > JR
- JR=
matrix.lduAddr().upperAddr()[0...mFaces-1]+1 - JC=
matrix.lduAddr().lowerAddr()[0...mFaces-1]+1 - AA=
matrix.upper()
- JR=
-
lower terms: JR > JC
- JR=
matrix.lduAddr().lowerAddr()[0...mFaces-1]+1 - JC=
matrix.lduAddr().upperAddr()[0...mFaces-1]+1 - AA=
matrix.lower()
- JR=
-
boundary term, only considering single processor problem here. There are 3 types of patch types:
- geometric (constraint) type.
- basic
- derived
In the following program, I assume there is not coupled interface such as processor patch or cyclic patch.
reference: Matrix coupling of different processors
Here is the code.
-
A c++ library
cnpyis used to generate npy or npz file for numpy.- site: https://github.com/rogersce/cnpy
- command:
git clone https://github.com/rogersce/cnpy.git cd cnpy mkdir build cd build cmake .. -DENABLE_STATIC=ON make make install -
dumpFvMatrix.H
#pragma once // added by CatDog #include<iostream> #include<fstream> #include<string> #include"cnpy.h" void dumpFvMatrix(string path, const fvScalarMatrix& EqnPtr) { const label nCells = EqnPtr.diag().size(); const label nFaces = EqnPtr.lower().size(); const scalar* const __restrict__ diagPtr = EqnPtr.diag().begin(); const label* const __restrict__ uPtr = EqnPtr.lduAddr().upperAddr().begin(); const label* const __restrict__ lPtr = EqnPtr.lduAddr().lowerAddr().begin(); const scalar* const __restrict__ upperPtr = EqnPtr.upper().begin(); const scalar* const __restrict__ lowerPtr = EqnPtr.lower().begin(); std::vector<scalar> AA(nCells+2*nFaces); std::vector<label> JR(nCells+2*nFaces); std::vector<label> JC(nCells+2*nFaces); // diag for(label cell=0;cell<nCells;cell++) { AA[cell]=diagPtr[cell]; JR[cell]=cell; JC[cell]=cell; } for(label face=0;face<nFaces;face++) { AA[face]=upperPtr[face]; JR[face]=lPtr[face]; JC[face]=uPtr[face]; } for(label face=0;face<nFaces;face++) { AA[face]=lowerPtr[face]; JR[face]=uPtr[face]; JC[face]=lPtr[face]; } cnpy::npz_save(path,"nCells",&nCells,{1},"w"); cnpy::npz_save(path,"nFaces",&nFaces,{1},"a"); cnpy::npz_save(path,"AA",&AA[0],{nCells+2*nFaces},"a"); cnpy::npz_save(path,"JR",&JR[0],{nCells+2*nFaces},"a"); cnpy::npz_save(path,"JC",&JC[0],{nCells+2*nFaces},"a"); return; }- in
myIcoFoam.C
// ... #include "fvCFD.H" #include "pisoControl.H" #include "dumpFvMatrix.H" // ... // Non-orthogonal pressure corrector loop while (piso.correctNonOrthogonal()) { // Pressure corrector fvScalarMatrix pEqn ( fvm::laplacian(rAU, p) == fvc::div(phiHbyA) ); pEqn.setReference(pRefCell, pRefValue); pEqn.solve(mesh.solver(p.select(piso.finalInnerIter()))); if (piso.finalNonOrthogonalIter()) { phi = phiHbyA - pEqn.flux(); } if (runTime.timeIndex()==2) { Info<< "TimeIndex = 2, output matrix pEqn"<<endl; dumpFvMatrix("/OF/OpenFOAM/-v1706/run/cavity/pEqn.npz",pEqn); } } // ...optionsfile:
EXE_INC = \ -I$(LIB_SRC)/finiteVolume/lnInclude \ -I$(LIB_SRC)/meshTools/lnInclude \ -I/usr/local/include EXE_LIBS = \ -lfiniteVolume \ -lmeshTools \ -Wl,-rpath -Wl,/usr/local/lib -lcnpy- python script to read the matrix and find
LU-SGS's effectiveness.
import numpy as np import scipy as sp from scipy.sparse import coo_matrix,tril,triu,diags from scipy.linalg import norm data=np.load('pEqn.npz') nCells=data["nCells"][0] nFaces=data["nFaces"][0] AA=data['AA'] JR=data['JR'] JC=data['JC'] A=coo_matrix((AA,(JR,JC)),shape=(nCells,nCells)) L=tril(A,-1) U=triu(A,1) D=diags(AA[0:nCells],0) Dinv=diags(1.0/AA[0:nCells],0) delta= L.dot(Dinv).dot(U) print "norm of error of LU-SGS: norm(L*Dinv*U)=", norm(delta.toarray())/norm(A.toarray()) # a Laplacian operator # proof of diagonally dominance print abs(np.abs((L+U).toarray()).sum(1)/np.abs(D.toarray()).sum(1)-1).max() # proof of symmetry print abs(L-U.T).sum() ## reference: Jisheng Kou, Yitian Li, A uniparametric LU-SGS method for systems of nonlinear equations, In Applied Mathematics and Computation, Volume 189, Issue 1, 2007, Pages 235-240, ISSN 0096-3003, f=lambda w:norm(((1-w)*(L+U)-w*w*L.dot(Dinv).dot(U)).toarray())/norm(A.toarray()) for w in np.linspace(0,1,100): print f(w)最后结果:
>>> >>> print "norm of error of LU-SGS: norm(L*Dinv*U)=", norm(delta.toarray())/norm(A.toarray()) relative norm of error of LU-SGS: norm(L*Dinv*U)= 0.141875980931 >>> ... # a Laplacian operator ... # proof of diagonally dominance ... print abs(np.abs((L+U).toarray()).sum(1)/np.abs(D.toarray()).sum(1)-1).max() 0.5 >>> # proof of symmetry ... print abs(L-U.T).sum() 0.0
